Viewing Jupyter notebooks at the command line
The Jupyter notebook is a literate programming environment that’s common in machine learning. The standard tools for working with notebooks are web applications, but sometimes you just want to view one at the command line. This comes up when you’re SSH’d into a training workstation and don’t want to set up port forwarding, start a Jupyter server, and open a browser just to glance at a notebook.
Other literate programming formats such as R Markdown and Pweave are easy to view at the command line. They use lightweight markup languages that don’t need special rendering to be legible, so you can usually just cat the file to the terminal.
Jupyter notebooks are different. A notebook is a structured JSON document conforming to a schema. Notebooks can contain images and other binary data, which, when viewed with a pager, will print unintelligible Base64 encoded data to your terminal. Characters such as " are backslash-escaped, making copying code from the output impossible.
Instead of printing the notebook file directly to the terminal, let’s convert it to another format first. nbconvert is a program to convert notebooks to rich formats, including HTML and PDF, as well as plain text such as Markdown. Markdown is readable as plain text in a terminal, so it’s a good choice of output format.
As an example, we’ll use a notebook that contains both prose and code cells taken from the JAX documentation.
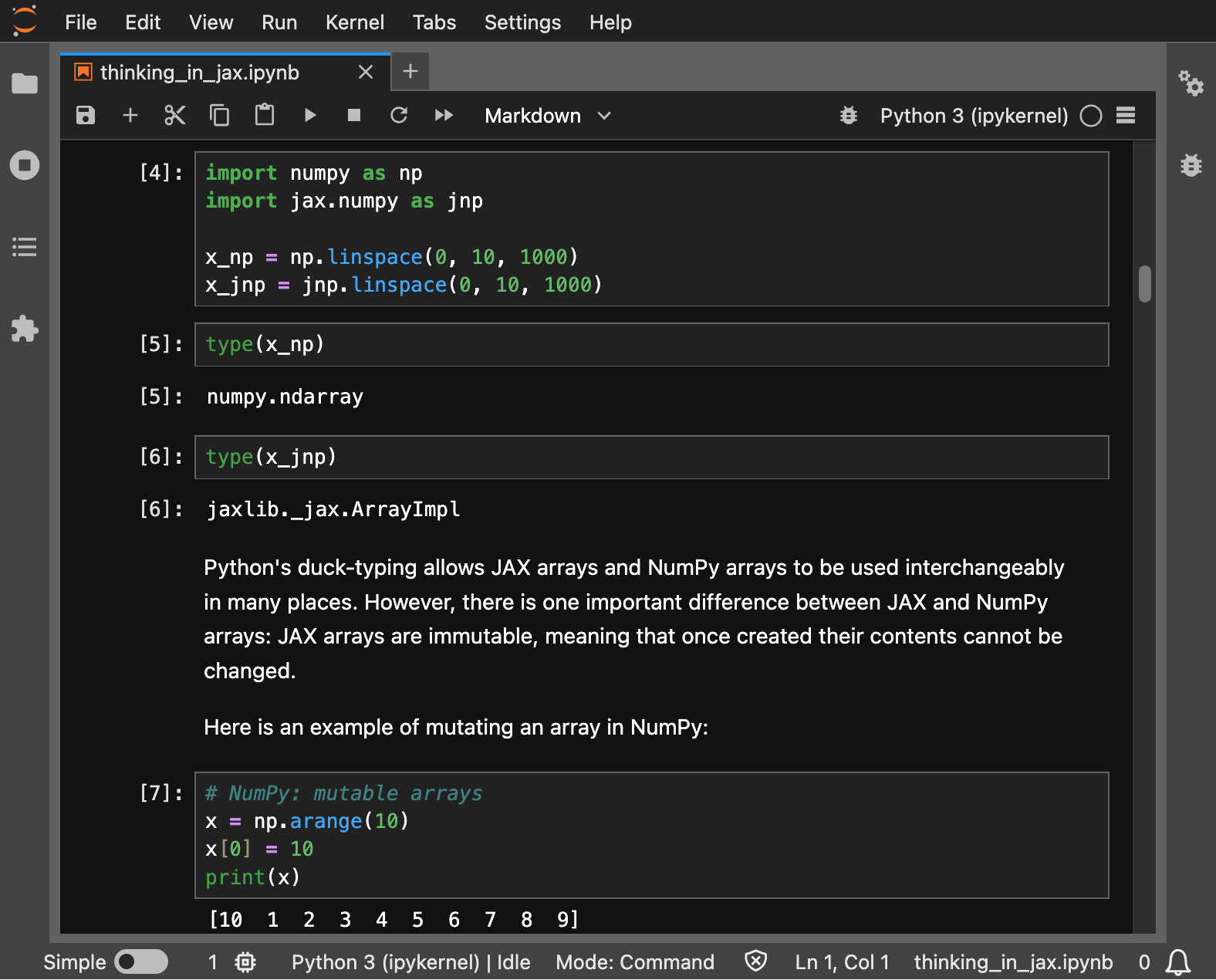
You can use jupyter nbconvert to convert the notebook to Markdown, printing the output to standard output, with:
jupyter nbconvert --stdout --to markdown thinking_in_jax.ipynb
nbconvert writes a header, [NbConvertApp] Converting notebook thinking_in_jax.ipynb to markdown, to standard error and the notebook body to standard output, so filter out the header by redirecting standard error:
jupyter nbconvert --stdout --to markdown thinking_in_jax.ipynb 2>/dev/null
This redirection syntax is for Bourne-like shells like bash and zsh. If you use an alternative like fish or Nushell, you’ll need its equivalent.
This is already more readable than the raw JSON, but we can improve the readability of code cells by applying syntax highlighting. We’ll pipe the output of nbconvert to pygmentize, a command line interface to the Pygments syntax highlighting library. Pygments is a dependency of nbconvert, so it should already be installed.
As we’re piping text to pygmentize on standard input, there’s no filename from which to determine the language of the input, so we need to specify it using the -l flag:
jupyter nbconvert --stdout --to markdown thinking_in_jax.ipynb 2>/dev/null |
pygmentize -l md
Let’s pipe the output of pygmentize to a pager like less to scroll through and search within the notebook:
jupyter nbconvert --stdout --to markdown thinking_in_jax.ipynb 2>/dev/null |
pygmentize -l md | less -R
The -R flag instructs less to output ANSI escape sequences in raw form to colour the output. If you run this command, you’ll get a legible rendering of the notebook with syntax highlighting. Thanks to less, you can scroll up and down through the notebook, and search within it by pressing / and typing a query.
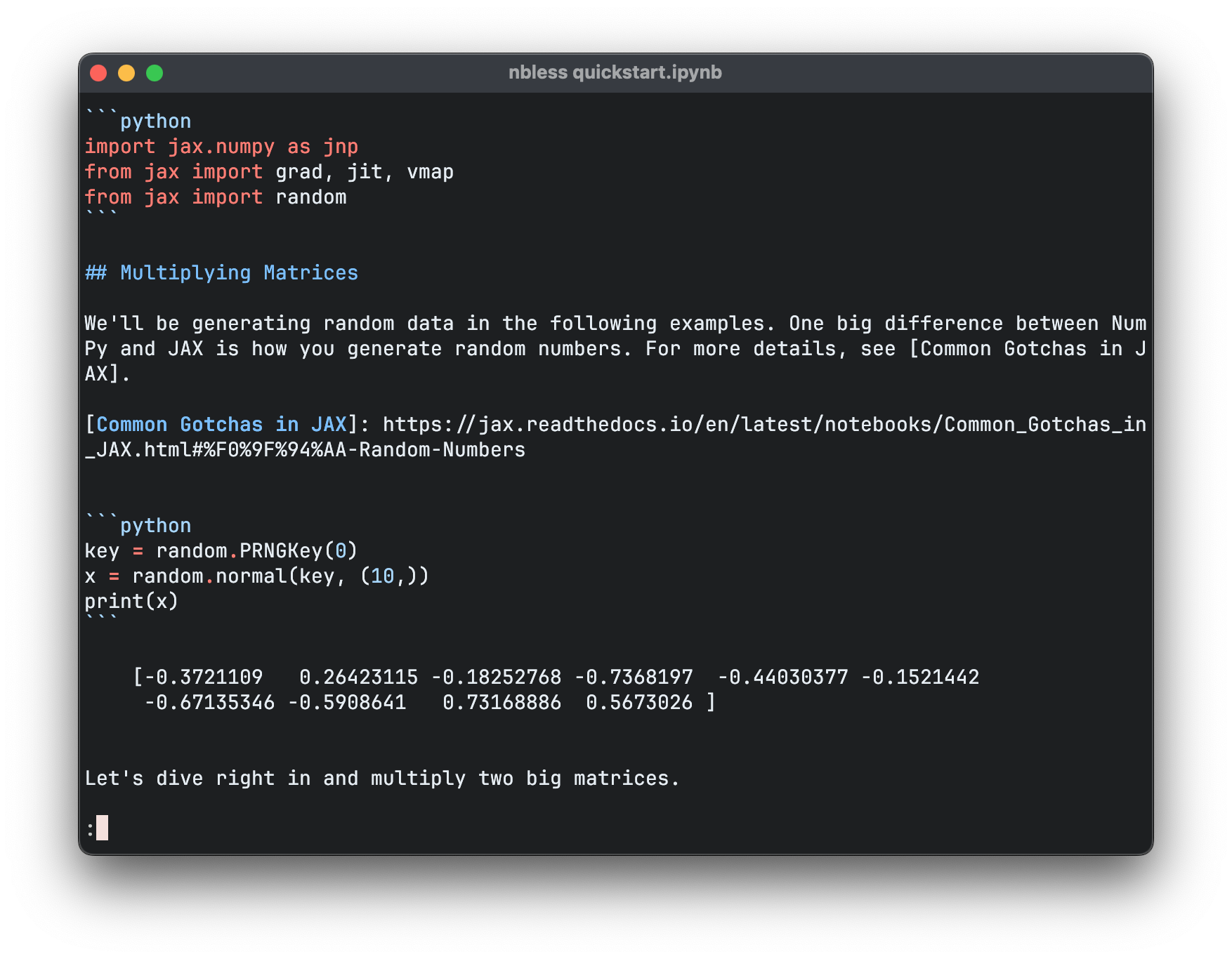
This is a longer command than is convenient to type for each use, so let’s create a shell script to encapsulate it, and also take the opportunity to add some input validation:
#!/usr/bin/env bash
set -euo pipefail
readonly EX_USAGE=64
readonly EX_NOINPUT=66
readonly EX_UNAVAILABLE=69
usage() {
echo "usage: $(basename "$0") [-h] NOTEBOOK"
echo
echo "Convert a Jupyter notebook to Markdown and view with less"
}
if [[ $# -ne 1 ]]; then
usage
exit $EX_USAGE
fi
if [[ "$1" = -h || "$1" = --help ]]; then
usage
exit
fi
if [[ ! -f "$1" ]]; then
echo "$(basename "$0"): $1: No such file"
exit $EX_NOINPUT
fi
if ! command -v jupyter >/dev/null; then
echo >&2 "$(basename "$0"): command not found: jupyter"
exit $EX_UNAVAILABLE
fi
jupyter nbconvert --stdout --to markdown "$1" 2>/dev/null |
pygmentize -l md | less -R
Save this as an executable script on your shell’s command search path, for example as nbless. Next time you want to view a notebook from the command line, run nbless notebook.ipynb.